Breaking down the microbiology world one bite at a time
An ancient chemical weapon against viral predators.
Viruses cannot disseminate their genetic material on their own. Crucial to their proliferation is the ability to infect a host organism and co-op host resources for viral transcription and replication. Many potent antiviral drugs used for clinical treatment of human viruses, such as herpes, HIV, or hepatitis C, target viral transcription and replication functions. Specifically, these therapeutics are small-molecule mimics of nucleotides, the building blocks of RNA and DNA, that are incorporated by viral polymerases during RNA or DNA synthesis. The mimics are like broken chain links, as their incorporation prevents the addition of more nucleotides by viral polymerases, thereby terminating polymerization. The consequence of this is inhibition of viral replication. In fact, this clinical strategy is analogous to an antiviral strategy already present in nature.
During viral infection, humans produce nucleotide chain-terminating molecules, and this ability relies on a protein called viperin. Viperin catalyzes the conversion of the ribonucleotide CTP (a component of RNA) to a modified molecule called ddhCTP. Like the clinical small-molecule mimics, ddhCTP inhibits viral transcription and replication, thus enabling broad antiviral activity against RNA and DNA viruses.
Like humans, prokaryotes, which include bacteria and archaea, must fend off viral predators. The “immune system” of prokaryotes compared to the human immune system is vastly different. Remarkably, however, Bernheim and colleagues recently discovered that the human capacity to synthesize nucleotide chain-terminators likely originated in prokaryotes. Their work was inspired by the observation that some bacteria and archaea (about 1%) encode homologs of human viperin. Here, the researchers showed for the first time that putative prokaryotic viperins, or pVips, defend against bacteria-infecting viruses called phages, via the production of chain-terminating nucleotides.
Given their discovery that some prokaryotes encode viperin homologs, the researchers hypothesized that these proteins, like human viperin, could function in antiviral defense. To test this, the researchers expressed pVips in the bacterial model organism Escherichia coli and challenged the E. coli with a diverse array of phages. They found that pVips could protect against or reduce infection. The enzymatic activity of the pVip was critical for this protection, as mutation of the key features required for catalysis abrogated the capacity to defend against phages. Surprisingly, in another repetition of this experiment, E. coli expressing the human viperin gene were also protected against phage infection. Thus, a human protein can combat a bacteria-specific virus. Together, their findings suggested that the function of prokaryotic pVips was similar to that of human viperin and, moreover, that the mechanism of defense is highly conserved among phylogenetically distant organisms.
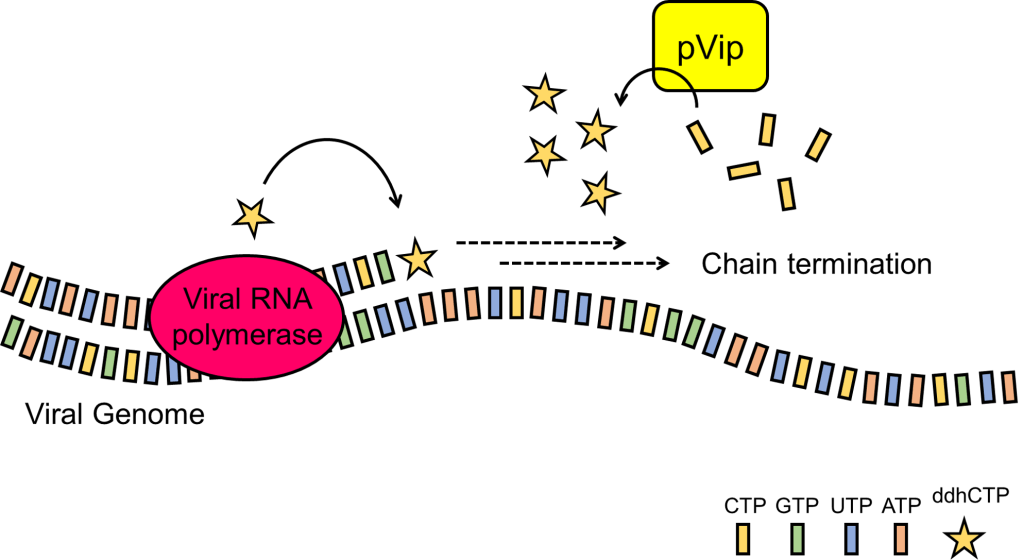
Key to the antiviral function of human viperin is the synthesis of ddhCTP. Thus, the researchers next goal was to demonstrate that prokaryotic pVips also produce this chemical weapon. Analysis of the small molecules harvested from E. coli expressing pVips revealed the presence of ddhCTP ribonucleotides. Additionally, some pVips could synthesize other chain-terminating molecules like ddhUTP or ddhGTP (derivatives of the RNA ribonucleotides UTP and GTP, respectively). Select pVips also had the capacity to produce multiple chain-terminating nucleotides, suggesting that these pVips are promiscuous and could harbor a diverse arsenal to protect against phage infection. They further showed that, in a test tube, purified pVip proteins converted ribonucleotides to their modified versions. Thus, the production of chain terminators like ddhCTP, ddhGTP, and ddhUTP is key to pVip antiviral function.
Finally, to understand the evolutionary origins of pVips, Bernheim and colleagues clustered pVips into groups based on DNA sequence similarity. Curiously, their analysis revealed that human viperins were not the most divergent from pVips encoded by bacteria and archaea. Rather, human viperins are positioned between a group dominated by archaeal species, and a group composed of bacteroides, a specific phylum of bacteria, with the human viperins’ closest relatives being the archaeal pVips. This finding suggested that eukaryotes (the delineation humans are a part of) likely acquired viperins from archaea as opposed to evolving pVips independently.
Bacteria deploy a diversity of strategies to combat predatory phages. Knowledge of bacterial defense strategies has provided a wealth of molecular tools for scientists. For example, proteins called restriction endonucleases, which bacteria use to cut invading viral genetic material, are widely used by molecular biologists to engineer DNA. Bacterial defense strategies could also inform clinical approaches to treating viral infection in humans. Interestingly, small-molecule chemical defense strategies are not widely observed in prokaryotes; pVips are one of the few reported examples. Perhaps ancient microbes harbor a host of antiviral small molecules that could be applied in a clinical setting. These, however, await further discovery.
Link to the original post: Bernheim A., Millman A., Ofir G., Meitav G., et al. Prokaryotic viperins produce diverse antiviral molecules. Nature. 2021. 589(7840): 120-124.
Featured image: Escherichia coli with phages from Wikicommons under the creative commons 4.0 licence