Breaking down the microbiology world one bite at a time
Microbes in cheese: competition or cooperation.
Microbial communities are everywhere in, on, and around us. DNA sequencing technology has enabled characterization of the composition, diversity, and complexity of microbial communities across ecosystems. However, given that much of our understanding of microbiology and molecular biology comes from studies using a genetically-tractable and easy-to-grow strain, such as the model bacterium Escherichia coli, many of our molecular tools are insufficient to investigate complex inter-species and inter-kingdom interactions. Thus, understanding the biology of multi-species communities found in nature requires a different approach. Specifically, many scientists have taken to developing simple model communities derived from fermented foods—like cheese. As a cheese ages, a biofilm, or surface attached community, of bacteria and fungi forms on its surface. This type of biofilm is commonly referred to as the cheese rind.
Researchers in the Dutton lab use cheese rind biofilms as an experimentally tractable system to study microbial community development and function [1, 2]. In their latest work, Dutton’s group, led by postdoctoral scholar Emily Pierce, applied their cheese rind model system to pinpoint specific genes in bacteria crucial to supporting growth in the presence of fungal partner. One of the species of bacteria they tested was E.coli to take advantage of the genetic resources available. The genes they discovered as important for growth in this context provided molecular clues to the nature of the bacterial-fungal interaction at play.
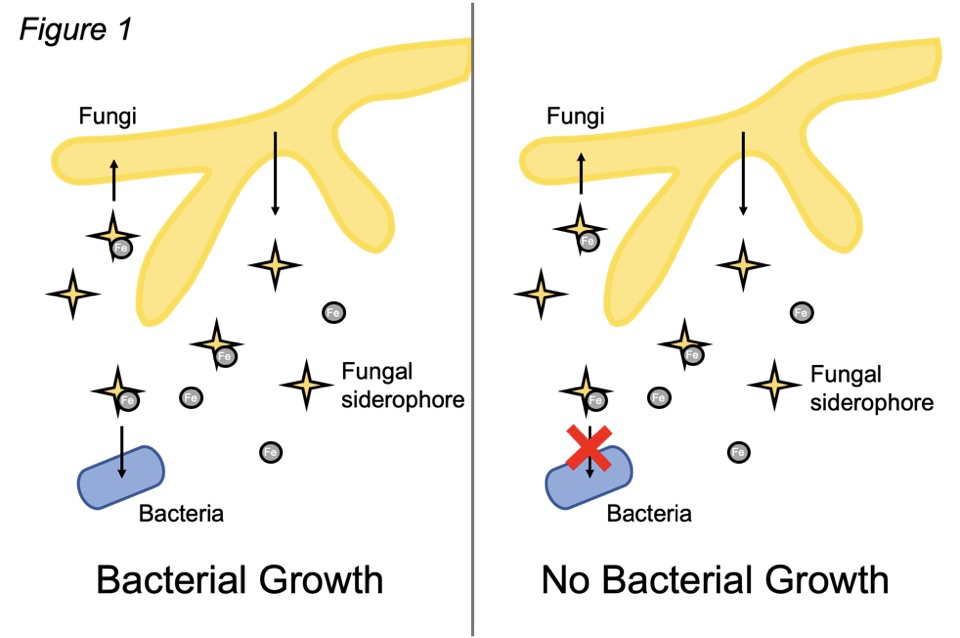
In iron-limited environments, microbials depend on siderophores, to scavenge for iron and support growth [2,3]. Dutton’s group discovered that, when grown alone, E. coli mutants that are defective in siderophore biosynthesis and uptake harbor a growth defect. However, when grown with a fungal partner, growth is rescued (Figure 1, left). This result suggested that, in the absence of its own siderophore, E. coli must leverage another source of siderophores. Indeed, E. coli encodes a separate siderophore transport system, which enables the capacity to uptake fungal-produced siderophores [4,5]. Consistent with this, fungi fail to rescue the growth of E. coli that possess mutations in uptake systems for both their own siderophores and the fungal siderophores (Figure 1, right). Thus, the researchers’ genetic analyses in E. coli uncovered that bacteria benefit from fungal siderophores to support their own growth (particularly in the absence of their own siderophores). Importantly, Dutton’s group observed that fungi commonly present in soil and our skin also alleviate the need for bacteria to produce their own siderophores, suggesting that the exchange of iron-scavengers is a common feature of bacterial-fungal partnerships in biological contexts beyond cheese rind communities.
The results of this study reinforce that bacterial-fungal partnerships are unquestionably complex, as the types of interactions varied among different combinations of bacterial and fungal partners. For example, their analyses suggest that only some fungal partners inhibit bacterial growth via the production of anti-microbials (Figure 2). Similarly, between the bacterial strains tested, the number of genes that are important for growth in this context, and the severity of these effects, per fungal partner varied significantly. Regardless, Pierce and colleagues were able to identify common themes that span various combinations of bacterial-fungal partnerships: competition for nutrients, like amino acids and biotin (a vitamin), and, as noted, the sharing of public goods like siderophores (Figure 2). The results of this study highlight that much remains to be discovered in the context of bacterial-fungal interactions, and inter-kingdom interactions more broadly. Progress towards the study of microbes in the context of a community, such as the cheese rind communities highlighted here, will be critical for understanding how microbial communities develop and function in natural settings.

WORKS CITED
1. Wolfe BE, Button JE, Santarelli M, Dutton RJ. Cheese rind communities provide tractable systems for in situ and in vitro studies of microbial diversity. Cell. 2014;158: 422–433. doi:10.1016/j.cell.2014.05.041
2. Pierce EC, Morin M, Little JC, Liu RB, Tannous J, Keller NP, et al. Bacterial–fungal interactions revealed by genome-wide analysis of bacterial mutant fitness. Nat Microbiol. 2021;6: 87–102. doi:10.1038/s41564-020-00800-z
3. Neilands JB. Siderophores: Structure and function of microbial iron transport compounds. J Biol Chem. 1995;270: 26723–26726. doi:10.1074/jbc.270.45.26723
4. Sauer M, Hantke K, Braun V. Ferric-coprogen receptor FhuE of Escherichia coli: Processing and sequence common to all TonB-dependent outer membrane receptor proteins. J Bacteriol. 1987;169: 2044–2049. doi:10.1128/jb.169.5.2044-2049.1987
5. Fecker L, Braun V. Cloning and expression of the fhu genes involved in iron(III)- hydroxamate uptake by Escherichia coli. J Bacteriol. 1983;156: 1301–1314. doi:10.1128/jb.156.3.1301-1314.1983



































































































