Breaking down the microbiology world one bite at a time
Bacterial Personality Uncovered With Single-Cells
Over the last decades interest in the behaviour of individual cells in mammals rapidly increased while bacteria were left in the dark. In 2009, Tang and his colleagues were the first to elucidate the functioning of individual cells in a developing mouse embryo by looking at the messenger RNA (abbreviated commonly as mRNA) content of individual cells. Herein, the field of single-cell transcriptomics was born. The mRNA is interesting as it is as it partially determines how a cell behaves. The mRNA is ‘read’ or ‘transcribed’ from the DNA code of a cell and subsequently translated into protein. Analyzing the transcribed mRNA content in the case of the individual cell is thus referred to in scientific terms, ‘single-cell transcriptomics’. It wasn’t until 2015, that Wang and colleagues demonstrated the potential of transcriptomics in individual bacterial cells, launching the field of bacterial single-cell transcriptomics. Unfortunately, despite the existence of a limited range of bacterial platforms, several technical constraints still make it difficult to discover individual bacterial behaviour. Below, we outline the tools and their use-case potential in this exciting new field of bacterial single-cell transcriptomics.
The phenomenon of single-cell behaviour
Today, bacteriomes are commonly studied at the level of microbial taxa or single species. Evidence suggests however, that for many bacterial species, this is short-sighted. To understand single-cell behaviour, consider a single colony of bacteria exposed to a similar environment, sharing an identical ancestry and genetic code. In this colony, individual bacterial cells, even though identical, exhibit different behaviour (see Fig. 1). More importantly, even in the absence of genetic variability, this behavioural heterogeneity helps the population as a whole exploit different nutritional niches or resist certain stressors. In that sense, individual cellular behaviour constitutes a type of ‘bacterial single-cell personality’. The mechanisms, whether stochastic or directional, driving variation at this level are, however, poorly understood to date.
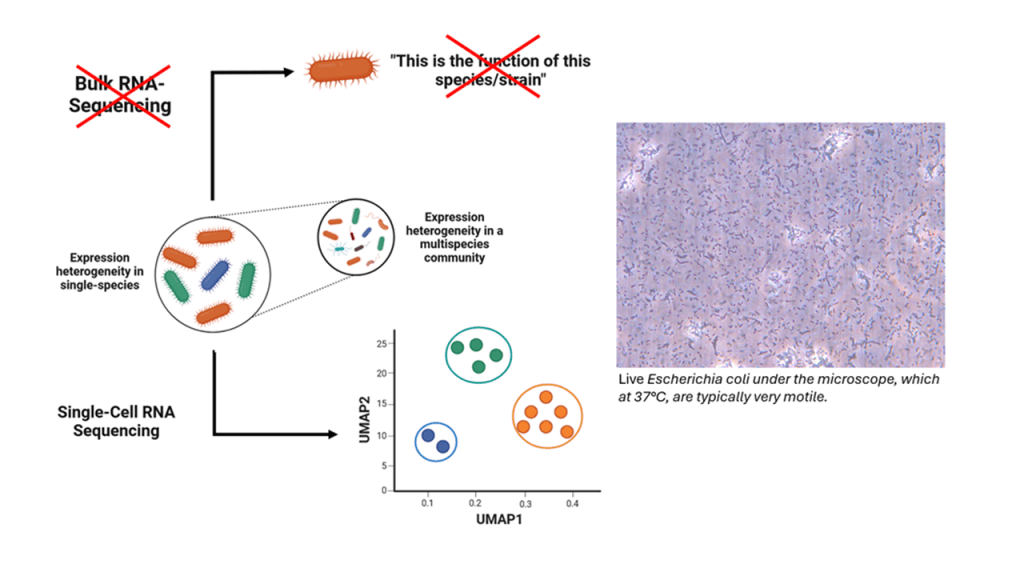
The principle behind revealing single-cell behaviour
Revealing what individuals are doing within a population of cells is not trivial. To determine what functions a single cell fulfills, identifying its expression profile or mRNA content is most meaningful. To assign a unique identity to a single cell, its mRNA content needs to be uniquely labelled before sequencing through a process called ‘barcoding’. In barcoding, unique DNA fragments are added to the mRNA in a cell-specific manner. As such, the short DNA fragments are directed in a manner that allows every cell to receive a unique molecular barcode.
High-throughput & low-throughput methods for assessing single-cell behaviour
All current bacterial single-cell RNA sequencing (also abbreviated as scRNA-Seq) platforms utilize a barcoding approach. Different strategies, however, exist to acquire uniquely barcoded cells, depending on the preferred throughput. In high-throughput scRNA-Seq platforms, acquiring a large quantity of cells is favoured, usually between 10 000 – 1 000 000 cells in a single run. In contrast, low-throughput scRNA-Seq platforms favour more extensive (or even full) coverage of the mRNA content of a single cell, but suffer from a lower throughput, such as 96 cells in the case of MATQ-Seq. Hence, prioritising throughput has the unfortunate side-effect of limiting the resolution at which an individual cell can be analyzed.
Fundamental strategies of state-of-the-art methods
Besides differing in focus, scRNA-Seq methods also vary in their method of sorting or isolating single cells to perform the aforementioned barcoding. Herein, three different strategies can be distinguished: plate-based methods, droplet-based methods, and fluorescence-based methods. Plate-based methods are the simplest, and also cheapest. In plate-based methods, reactions in which DNA fragments (‘barcodes’) are added to the mRNA happen inside a permeabilized cell, where the scientist makes the membrane semi-permeable to allow barcodes to enter the cell. Subsequently, the cells are sorted in a typical laboratory well plate in different wells, each containing a unique barcode which is attached to the mRNA. Through pooling the barcoded cells and repeating this procedure, the process can be scaled up. In this manner, more cells can be barcoded as each cell has a random chance of ending up in each barcode well. This is ‘combinatorial barcoding’, which is the fundamental principle of methods such as MicroSPLiT and PETRI-Seq.
In droplet-based methods, in contrast, a single cell is encapsulated by a specialized microfluidics device inside a droplet. A droplet is nothing more than a small aqueous compartment surrounded by a lipid membrane. As a cell gets encapsulated in a droplet, scientists release the mRNA content of that cell inside the droplet after which barcodes are added to the mRNA. As droplet-based methods require a specialized device and expertise, they are generally considered more difficult. Examples of droplet-based methods (often used in combination with a plate-based approach) are BacDrop, smRandom-Seq, and ProBac-Seq.
Lastly, fluorescence-based methods such as par-SeqFISH rely on the principle of fluorescence-in-situ hybridization (FISH). Although they provide additional information on the spatial location of bacterial cells within a tissue, they are limited to how many fluorophores can be resolved and designed for your bacterial species of interest. Hence, researchers can maximally assess only a few cell functions.
Outlook on the field of bacterial single-cell transcriptomics
Most methods adapt eukaryotic methodology to a challenging bacterial system. Bacteria lack a tail of adenines in their mRNA which is typically used to capture eukaryotic mRNA. Moreover, bacteria contain very little RNA, and of this RNA, over 90% is ribosomal RNA, whose expression is not useful for separating single-cells. Lastly, the bacterial kingdom is incredibly diverse, leading to potential sampling bias, and issues acquiring permeabilized cells. Thus, as bacterial single-cell transcriptomics is expensive, and requires special equipment and expertise, its adoption has been limited. However, with adequate technical cross-comparison between methods, future adoption will require less optimization and be less expensive. Moreover, as the bacterial community realizes the importance of ‘single-cell personality’, I expect its influence to pick up pace.
Written by Camiel Corten
Link to the original post: Andrew W. Pountain, Itai Yanai, Dissecting microbial communities with single-cell transcriptome analysis. Science 389, eadp6252(2025). DOI:10.1126/science.adp6252
Featured image: Artist’s interpretation of the observation of a green oval bacterium and bacterial behavior at the single-cell level. This work was made by Belgian illustrator Kyran Dewit



































































































