Breaking down the microbiology world one bite at a time
A simple Switch from Friend to Foe.
The world around us is teeming with microorganisms, from the deep depths of the ocean to our own gut. Microbes are commonly labeled as infectious when, in reality, only a small portion is classified as pathogens. Many microbes live happily without causing any harm to us or other forms of life; they can even benefit their host by providing them with protection or essential nutrients.
Then what makes a pathogen a pathogen?
The key to a microbe’s ability to infect a given host lies within its genome. Pathogens harbor a collection of tools encoded by its genes to attack a host or avoid the host’s immune defenses. Commensal microbes generally don’t possess these genes, or if they do, they don’t express them. Other bacteria, known as opportunistic pathogens, can seamlessly switch from harmless friends to infectious foes.
Many scientists are exploring this opportunistic switch in a bacterium called Staphylococcus epidermidis. S. epidermidis is one of the many microbes normally found on our skin. While typically benign, this bacterium can be a problematic pathogen in hospitals and clinics. S. epidermidis often grows on implanted medical devices and occasionally even infects the blood, leading to life-threatening diseases like sepsis. It has also acquired resistance to a number of antibiotics that makes treatment a continuous challenge. Interestingly, S. epidermidis doesn’t have many genes that help attack its host directly. It instead relies on genetic tools to sneak past the host’s immune response, invade, and establish residence in the host.
How does this microbe go from being harmless on our skin to causing life-threatening infections?
Previous research by Dr. Laura Cau and others found that some S. epidermidis strains naturally produce more proteases, known as EcpA, than others that lead to skin damage and inflammation. To learn more about EcpA in S. epidermidis, check out The skin’s “frenemy”. In a recent study, a collaborative group of researchers from Germany, Denmark, and China found that strains of S. epidermidis have switched from a commensal to pathogenic lifestyle with the help of a complex sugar molecule known as RboP wall teichoic acid (WTA). WTA is fairly common in bacteria and squeezes itself between peptidoglycan on the bacterial cell wall (Figure 1).

It likely plays critical roles in how bacteria maintain their shape, control cell processes, and interact with their hosts [2]. All strains of S. epidermidis possess WTA, but only pathogenic strains have RboP-WTA. The researchers demonstrate that RboP-WTA changes how infectious S. epidermidis colonizes its host and allows it to share some of its genetic tools with Staphylococcus aureus, a pathogenic relative to S. epidermidis, to enhance its new virulent lifestyle.
The team of researchers used clinical samples from S. epidermidis-infected patients to understand key differences between harmless and infectious strains. They found that pathogenic S. epidermidis strains possess two types of WTA: GroP-WTA and RboP-WTA. Non-pathogenic S. epidermidis strains, on the other hand, only have GroP-WTA (Figure 2).

S. aureus also expresses RboP-WTA, which suggests that RboP-WTA enables a pathogenic lifestyle for S. epidermidis. Further research revealed that RboP-WTA changes how the bacterium interacts with and colonizes its host. According to the study, S. epidermidis expressing RboP-WTA easily infects the host’s bloodstream and loses its ability to colonize host epithelium like the commensal strains.
Bacteria often team up to survive and evolve by exchanging valuable genes. Since S. aureus and pathogenic S. epidermidis both express RboP-WTA, the authors wanted to see if the two species are capable of swapping DNA. They found that only S. epidermidis strains with RboP-WTA harbor a cluster of WTA-related genes that allow the bacteria to exchange DNA with S. aureus. Commensal S. epidermidis lacks both the gene cluster and the ability to trade genetic information with their S. aureus relatives. The authors hypothesize that the pathogen’s ability to exchange DNA with S. aureus provides S. epidermidis the perfect opportunity to learn how to better attack its host and resist common antibiotics.
These findings may open the door to the production of Staphylococcal vaccines that target RboP-WTA without disturbing our normal microflora, which is a huge step forward in the fight against antibiotic-resistant bacterial pathogens.
If you want to know more about this species, check Danny Ward #52Species about Staphylococcys epidermidis!
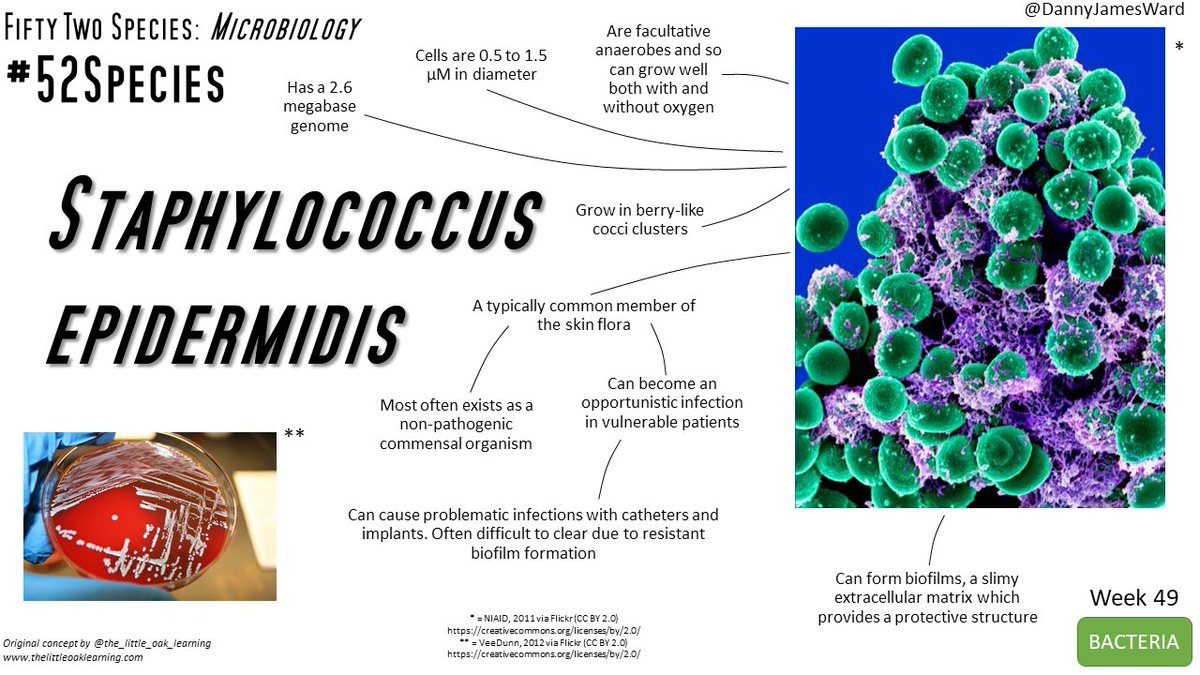
Originally tweeted by Danny Ward (@DannyJamesWard) on June 28, 2021.
Link to original article: Du, X., Larsen, J., Li, M. et al. Staphylococcus epidermidis clones express Staphylococcus aureus-type wall teichoic acid to shift from a commensal to pathogen lifestyle. Nat Microbiol 6, 757–768 (2021).
Other references:
- Cau, Laura et al. “Staphylococcus epidermidis protease EcpA can be a deleterious component of the skin microbiome in atopic dermatitis.” The Journal of Allergy and Clinical Immunology vol. 147 (2021): P955-966. doi:10.1016/j.jaci.2020.06.024
- Brown, Stephanie et al. “Wall teichoic acids of gram-positive bacteria.” Annual review of microbiology vol. 67 (2013): 313-36. doi:10.1146/annurev-micro-092412-155620
Featured image: Created with BioRender.
Written by Julie Starkey



































































































