Breaking down the microbiology world one bite at a time
Phages reinterpret the genetic code
Crucial to all living cells is the ability to read the instructions stored in DNA on how to make proteins. This flow of information is driven by two processes: transcription and translation. During transcription, DNA is copied into a messenger RNA (mRNA) sequence. Next, during translation, the mRNA sequence is used as a template to build a sequence of amino acids in protein synthesis. In this case, three mRNA nucleotides, called a codon, specify a particular amino acid. Some codons, i.e., “stop codons” instruct the cell’s translation machinery to terminate the addition of more amino acids. This set of rules that cells must follow to translate the information stored in their DNA into proteins is called the genetic code, and the genetic code is highly conserved across life.

That being said, there are rule-breakers. One such group of rule-breakers are bacteria-infecting viruses, called phages. Phages are obligate parasites, dependent on their host bacterial cell for resources to persist. For instance, many viruses do not encode their own protein translation machinery and therefore must hijack their hosts’ capabilities to build viral proteins. Curiously, recent investigations have revealed that many phages have “recoded” their stop codons and reassigned them to amino acids. Scientists are now curious as to why some phages have evolved codes that are incompatible with their host bacterial translation systems.
In a recent study, Borges et al. took a computational approach to characterize phages with recoded stop codons. The goal of their work was to garner insight into the mechanism of recoding, and the potential function of recoding with respect to phage biology. They discovered a widespread use of recoded stop codons across multiple families, or clades, of phages. Six of the clades they newly discovered, which they named Garnet, Amethyst, Jade, Sapphire, Agate, and Topaz.
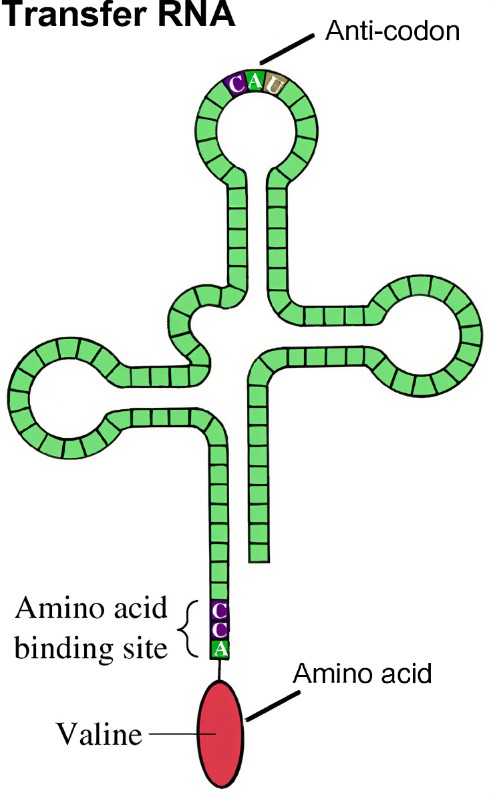
The researchers identify multiple ways phages could possibly reassign stop codons to amino acids. For example, they found that many of the phages encode molecules called suppressor transfer RNAs. In general, transfer RNAs (tRNAs) are a key component of protein translation as they are responsible for delivering the amino acid, corresponding exactly to the codon specified, to the translation machinery. When a stop codon is encountered, no tRNA delivers an amino acid, and protein synthesis is terminated. Thus, phage-encoded suppressor tRNAs, unlike “typical” tRNAs, are capable of recognizing the standard stop codons and inserting another amino acid.
Interestingly, not all genes within a single phage genome are recoded. Most commonly, alternative code is employed in defined regions of the phage genome, such as the structural and lysis genes. Expression of structural and lysis genes enables the phage to package its genetic material, kill its host cell, and then proceed to infect the next host cell. These sets of genes are often precisely regulated in a phage, as they drive a significant lifestyle switch from dormancy in the host cell, to cell-death and infection of a new host. By contrast, genes that enable the phage to integrate in the host genome and establish dormancy predominantly use standard genetic code. Similarly, genes that encode the suppressor tRNAs also tend to rely on the standard code.
These findings suggested to the researchers that genetic recoding could play a role in regulating the decision to switch to phage lysis. The model is that when a phage infects a host, the phage produces standard-code compatible proteins (using the bacterial host machinery). In some phages, the proteins produced at this step establish dormancy within the host cell. Next, phage molecules that are “code changers” are produced, such as the suppressor tRNAs. Consequently, recoded gene products that were initially not made, are now synthesized triggering host-cell lysis and phage dissemination.
The results of this work highlight the possibility of genetic recoding as an additional way phages control their lifestyle decision-making. Going forward, research will need to focus on experimental validation of the model proposed in this work. For instance, the researchers did not identify suppressor tRNAs in all phages that use alternative coding. One possibility is that, in the case of two phages co-infecting one bacterial host, phages that don’t harbor suppressor tRNAs could exploit the suppressor tRNAs produced by the co-inhabiting phage. Another possibility is that there are additional mechanisms of recoding that have yet to be discovered. Similarly, what controls the expression of code changers in these phages is unknown, and answering this question could further our understanding of the diverse inputs phages use to regulate their biology.
Link to the original post: Borges, A.L., Lou, Y.C., Sachdeva, R. et al. Widespread stop-codon recoding in bacteriophages may regulate translation of lytic genes. Nat Microbiol 7, 918–927 (2022).
Featured image: bacteria-phage – Bing images