
Breaking down the microbiology world one bite at a time
Evolution of the human pathogenic lifestyle in fungi
It is said that of the billion human infections caused by fungal pathogens every year, over 300 million people are afflicted with a serious fungal infection with more than 1.6 million deaths worldwide. Furthermore, only 30 out of 600 species of the noted fungi are known to infect humans regularly.
Despite their obvious presence in society and enormous annual global burden, fungi, unlike viral and bacterial infections, have been (severely) undermined and understudied. Arturo Casadevall in a workshop (2011) reinstated this by stating that less than 10% of 1.5 million estimated fungi have been formally described. In the same workshop, David Relman voiced out that the blindspot in regards to the distribution of fungal biogeography reflected on our “fairly poor situational awareness”.
Antonis Rokas (2022), while discussing the evolutionary aspects of human fungal pathogenicity, suggests that the inherent opportunistic nature of fungi is the reason these pathogens have evaded attention. This is why every fungus can be a threat as a potential pathogen.
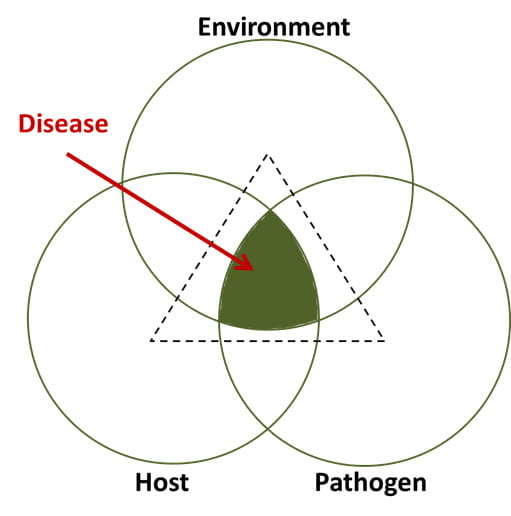
What is this “fungal pathogenicity” you ask? Well, Antonis defines it as “the outcome of complex interactions between pathogens, human hosts and their environment”.
As observed over the past few decades, fungal pathogens have displayed many unique traits and genetic elements that have bloomed their virulence to survive and thrive in humans. Evolved advantages of virulence are certainly not helpful as determinants to disease nor particularly unique but are assumed to have emerged for the survival of fungi independent of the human infection route. Nevertheless, these advantages are still rampant in ensuring their growth and survival within the host environment.
This is why, he states, “understanding fungal virulence requires understanding of the natural history, ecology and adaptations of fungi that facilitate their success in their natural environments.”
Many major fungal pathogens are assumed to evolve repeatedly and independently from non-pathogens across some major lineages, and this is often represented as a concept of convergent evolution, exemplified by the dimorphic fungi (C. albicans, Histoplasma, Blastomyces, and C. neoformans), which can switch from their mycelial filamentous state to yeast- like state.
Human pathogenic fungi differ greatly in their adaptation strategies to pathogenic lifestyles. Repeated evolution suggests that these pathogens may have developed certain pre-adapted traits that enable them to infect humans. Aspergillus, associated with the infamous aspergillosis, has evolved multiple times through A. fumigatus and A.flavus. Budding yeasts have evolved at least five times (!) independently in Candida glabrata, Candida albicans, and the emerging pathogen Candida auris, which causes candidiasis. Herein, pathogenicity has been observed to evolve drastically, burdening the health industry.

Image source: https://microbeonline.com/mycosis-etiology-types-and-classification/
Genes and traits that contribute to various forms of pathogenic behavior occur at the level of lineages, species and even strains. Strain level is interesting in two major ways: from the evolutionary point of view and clinical relevance. Understanding the former helps us to appreciate the underlying genetics in the organisms, while focussing on the clinical relevance helps us to understand the associated virulence and antifungal drug resistance profile. Strains of C. albicans and A. fumigatus in particular have been noted for their extensive genomic and phenotypic heterogeneity with respect to virulence and drug resistance profile.
Varying genes influencing pathogenic variation can be either through major changes which influence huge parts or even the entire genome, or small-scale changes affecting an individual genomic region. These large changes can arise from events like hybridization, variation in ploidy and/or transposon movement. On the other hand, small changes include gene duplication, gene loss, horizontal gene transfer, change in the copy number, and so on. This obviously gives rise to further variations in mechanisms along with diversifying strategies that encourage fungi to survive and flourish within the human host system.

The ability to colonize new ecological niches also highlights their versatile nature. Once fungi associate with humans, they adapt themselves by diverging from their natural phenotype. These fungal infections within human organisms could arise as a result of zoonotic outbreaks or through spores in the environment.
The ecological traits encouraging human-fungal pathogenesis include biogeographical distribution, tolerance at normal body temperature (thermotolerance), adaptability to varying oxygen levels, evasion from predation, and so on. These factors differentiate pathogens from their non-virulent peers, and help figure out the drive for evolving pathogenicity.
A noteworthy example of evolved pathogenicity from ecological traits has been illustrated by Patrícia Albuquerque et al (2019). According to them, repeated exposure of Paracoccidioides to predatory amoebae increased the ability of these fungi to infect mice and survive mammalian macrophages. This implies that the fungi have found a suitable way to evade a major protective strategy (macrophages) in our human immune system and infecting mice corresponds to the pathogenic nature of our culprit. A similar trend was observed in Cryptococcus, which, upon its exposure to amoeba, displayed pseudohyphal growth rather than yeast-like (unicellular form) followed by resistance to macrophages at the cost of reducing its virulence. These two examples show that upon repeated exposure to natural predators, fungi adapt themselves in novel interesting ways – at times, by compromising their advantageous traits (such as virulence) or even by changing their forms – to be able to thrive within the host environment.
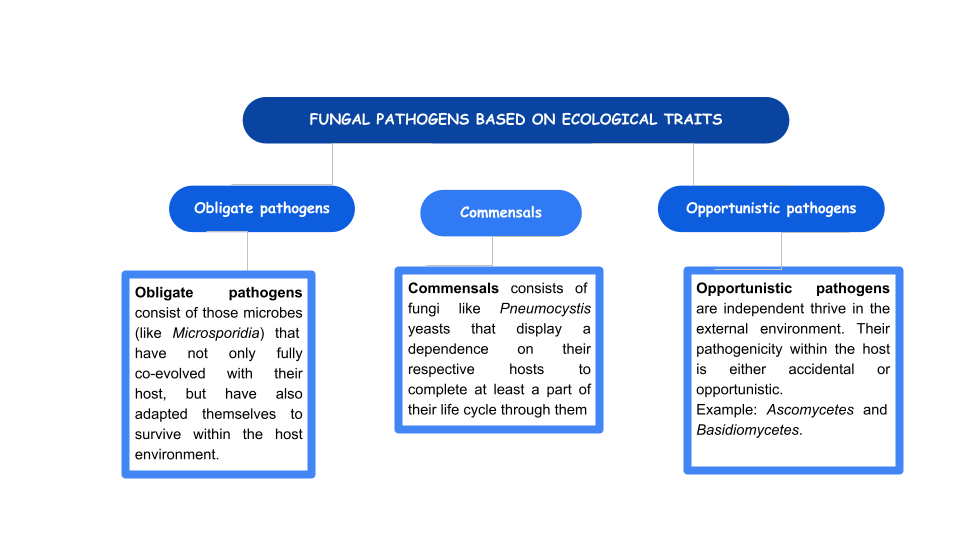
Incidentally, fungal genotypic and phenotypic variations can now be linked through various analytical techniques such as reverse ecology, phylogenetic methods and genome-wide association studies. Virulent traits can also be dissected by genetic and molecular tools such as genome sequencing.
But, we still do not know to what extent various observed phenotypes have a genetic basis, and this poses a challenge in targeting metabolic pathways to develop fungal vaccines and antifungal drugs. We have also yet to explore the natural lifestyle of these fungi and how the human host environment parallels the outer environment wherein these organisms thrive equally well. In short, we still have a lot to glean about the ecology and natural history of fungi!
And by now, if you still are wondering: why bother with the ecology of the world of fungi? Here’s why we need to: understanding their natural history and evolutionary ecology help us understand how disease-relevant traits have evolved independently in a repeated manner over time. Detecting and understanding those infection traits and the underlying mechanisms enabling such (devastating) pathogenicity can allow us to strategize ways to predict the emergence of fungal pathogens and develop drugs to mitigate the disastrous effects of fungal infections, as well as combat causal pathogens. Exploring the world of fungi and deciphering their sneaky routes of virulence is truly something worth figuring out, don’t you agree?
Link to the original post: Rokas, A. Evolution of the human pathogenic lifestyle in fungi. Nat Microbiol 7, 607–619 (2022). https://doi.org/10.1038/s41564-022-01112-0
Featured image: Created by the author using Biorender.com



































































































