Breaking down the microbiology world one bite at a time
When a curse becomes a blessing
Think of a cell as a thick pudding made out of proteins, if you are thinking of eukaryotic cells, lipidic structures that produce and use even more proteins will be enclosed within a bilipid layer with even more proteins embedded. For a cell, proteins are everything, they take care of DNA replication, transcription, protein production, energy harvesting and nutrient transport. Therefore keeping your proteins neat, well folded and functional is key to being alive. In order to ensure that, cells may use as much as half of their energy to keep a protein housekeeping system. This system is in charge of avoiding the production of protein aggregates that may interfere with the crowded intracellular environment.
When things go wrong, and the protein housekeeping system is not efficient, proteins can produce fibrillar forms called amyloid that tend to aggregate and at some point start to precipitate inside the cells. The reason behind the production of amyloid is not clear in all cases. But they could be due to chemicals like heavy metals, a mutation in proteins that make them less stable or degeneration of the protein housekeeping system due to aging. In any case, cell degeneration and eventually cell death are certain if aggregates are not cleared out.
The production of such protein aggregates is the cause of human illnesses associated with aging like Alzheimer’s disease and Parkinson’s disease. On the lookout for a cure to these and other amyloid-related diseases, we’ve been gathering over the years a lot of important information about what triggers protein aggregation, and now, some of that knowledge is being used to kill antibiotic-resistant bacteria.
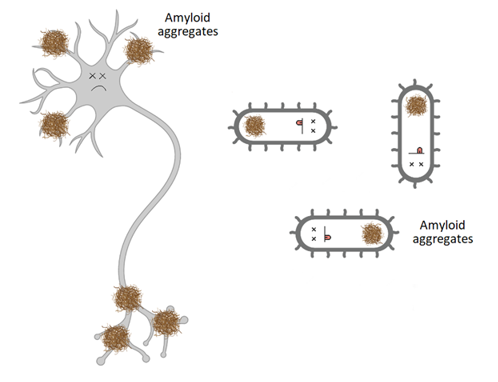
How is this done? First, you need to find regions prone to aggregation in the proteins of the bacteria you would like to kill. Later a small peptide with a similar sequence as these aggregation-prone regions is synthesised and pimped to help bacterial uptake. Lastly, the bacteria is treated with the peptide, where after uptake by the cell, the peptides will promote the aggregation of otherwise soluble proteins and would start a fast reaction where the proteins will form amyloid-like aggregates.
Then in the lab, we can kill bacteria by producing amyloid aggregation like in Alzheimer’s disease, but would it work in a real infection? Yes! This approach was used by Khodaparast and colleagues to treat a mouse model for bladder infection with really nice results, indeed it turns out that the peptide treatment administered intravenously is even more efficient in killing bacteria than antibiotics.
The mechanism behind these small peptides killing power is their design; each peptide can interact and produce the aggregation of more than one protein. Therefore by producing aggregation of multiple proteins with different functions the survival of the cell is jeopardised. This coupled with the accumulation of protein aggregates becomes an energetic burden. Later on, in the attempt to clear out protein aggregates, exhaustion of certain parts of the protein housekeeping system makes all the processes inefficient and triggers even more protein aggregation. Leading to total cell failure and death. This cascade of events makes it really difficult for a bacterial cell to build up mechanisms against the peptides, so far, bacteria cannot fight back and produce resistance to this kind of treatment, which is very promising for future therapeutics.
Even though more studies are needed before this technology can reach clinical trials, it is interesting to see that our knowledge from other research fields can be used to build up new antimicrobial tools like this one, that perhaps, can save us from the antibiotic apocalypse.
Link to the original post: Khodaparast L, Wu G, Khodaparast L, Schmidt BZ, Rousseau F, Schymkowitz J. Bacterial Protein Homeostasis Disruption as a Therapeutic Intervention. Front Mol Biosci. 2021 Jun 2;8:681855. doi: 10.3389/fmolb.2021.681855.
Featured image: Amyloid aggregates can be used to kill bacteria (created by author with Biorender)