
Breaking down the microbiology world one bite at a time
Prokaryotic origins of science’s biggest gene editing technology.
CRISPR is today’s most well-recognized gene-editing technology. It is faster, cheaper, and more efficient than any competing technologies. Scientists established that CRISPR can be used to edit the genomes of many plants and animals, including humans [1]. So-called “gene therapy” could potentially be used to treat human genetic diseases and reverse aging. For instance, Chinese scientists recently used CRISPR to edit the CCR5 gene of two twin babies [2]. CCR5 enables HIV infection, thus altering this gene could protect against viral infection. This example describes the applications of CRISPR in a novel context. But, what is CRISPR’s original context? While recognized for its use in human cells, this genetic tool is adapted from an antiviral mechanism found in prokaryotes.
CRISPR is the adaptive “immune system” of prokaryotes. Bacteria and archaea are frequently challenged by viral predators called phages. To protect themselves, prokaryotes use a variety of antiviral defenses, including CRISPR: Clustered Regularly Interspaced Short Palindromic Repeats. A specific region within the prokaryotic genome, called the CRISPR locus, contains multiple copies of the same sequences. These repeat sequences alternate with variable sequences known as spacers (Figure 1A) [3]. The key feature of spacer sequences is that they match sequences found in infectious phage predators and other foreign DNA elements. Thus, analogous to our own adaptive immune system, CRISPR spacers encode the molecular “memory” of previous viral attacks. The advantage of this is that when bacteria and archaea are challenged by a phage they have previously encountered, they can rapidly recognize and destroy the foreign genetic material, thus avoiding infection.

The presence of CRISPR sequences was first reported by Francisco Mojica in 1993 [1]. However, the biological function of this particular region of the genome was not fully understood. About two decades later, scientists working at a Copenhagen-based dairy company called Danisco determined the role of CRISPR in the bacterial adaptive immune response against phages [3]. Importantly, dairy industries are highly invested in understanding mechanisms of phage resistance since the bacterial cultures that are crucial to dairy fermentation are often susceptible to phage attack. The consequence of this is spoilage of the dairy products. In a key study, Danisco’s Rodolphe Barrangou and Philippe Horvath demonstrated experimentally, for the first time, that CRISPR sequences immunize bacteria against phages harboring homologous, or matching, genomic sequences [3].
For their investigation, Barrangou and Horvath selected the species Streptococcus thermophilus, a lactic acid-producing bacteria used in dairy fermentation, as a model to study CRISPR-dependent phage resistance. Analysis of the CRISPR loci of various phage-sensitive and phage-resistant S. thermophilus strains revealed that phage-resistant strains had more spacer sequences, suggesting that the spacers could be key to immunity [3].
How does S. thermophilus acquire spacers in its genome? Their work demonstrated that CRISPR loci are dynamic and rapidly change during phage exposure [3]. Specifically, the researchers exposed a virulent phage isolated from yogurt samples, called phage 858, to a vulnerable S. thermophilus strain. This allowed them to generate bacterial mutants that acquired immunity to phage 858. Analysis of the CRISPR DNA content of surviving bacteria consistently revealed the insertion of additional spacers into their genomes. Importantly, the researchers observed that the new spacer sequences corresponded exactly to the sequences found in the phage used for the infection. Additionally, bacteria that possessed any mismatches in their spacer sequence compared to the phage sequence were more vulnerable to phage infection. Together, these results indicated that sequence similarity between the phage and spacer is crucial for antiviral defense.
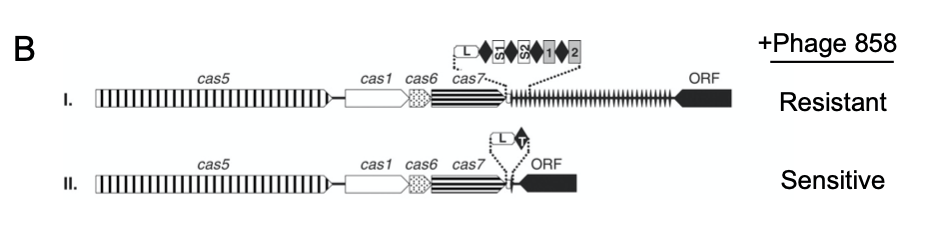
Given their findings, the researchers hypothesized that the information encoded by CRISPR spacers provides phage resistance. To test this, they altered the CRISPR loci of S. thermophilus strains and determined the effects on phage resistance (Figure 1B) [3]. First, the researchers focused on two spacer sequences, called S1 and S2, which they learned corresponded to sequences in phage 858. They introduced S1 and S2 into the CRISPR locus of an S. thermophilus strain sensitive to phage 858, and observed that the strain became resistant (Figure 1B, I). Second, they removed spacers from the CRISPR locus of an S. thermophilus strain that is immune to phage 858. Consequently, they observed that this strain could now be infected by phage 858 (Figure 1B, II). Thus, the presence of the spacers drives highly-specific immunity [3].
Spacers are not the only components necessary for CRISPR-based immunity. Adjacent to the CRISPR loci are a set of genes called cas genes that are also required for antiviral defense (Figure 1A and 1B) [1,3]. Since Barrangou and Horvath’s breakthrough findings, seminal studies have delineated the specific roles of cas genes in bacterial adaptive immunity [1]. This collective understanding of the underlying biology of CRISPR has allowed scientists to repurpose its core components for the development of new possibilities in the field of gene editing, thus launching an explosion of biotechnology applications relevant for human health and disease treatment.
Literature Cited
- Lander, E.S. (2016). The Heroes of CRISPR. Cell 164, 18-28.
- Raposo, V.L. (2019). The First Chinese Edited Babies: A Leap of Faith in Science. JBRA Assist Reprod. 23, 197-199.
- Barrangou, R., Fremaux, C., Deveau, H., Richards, M., Boyaval, P., Moineau, S., Romero, D.A., and Horvath, P. (2007). CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712.
Featured image: https://i1.wp.com/foodcrumbles.com/wp-content/uploads/2018/07/Home-made-yogurt-1-2.jpg?fit=750%2C500&ssl=1



































































































