BACTERIA, BIOFILMS AND FORENSIC PMSI

Breaking down the microbiology world one bite at a time
Bacteria, Biofilms and forensic PMSI
Did you know that microbes are applied in forensics to demystify unsolved murders? In fact, their presence in a biofilm on a carcass or a corpse can actually help solve “death-mysteries”. Let’s dive in.
Postmortem Submersion Interval Estimation (PMSI) refers to the time duration starting when a dead body enters a water body until its recovery. This estimation infers the time and location from where the body fell within the water. On extrapolation, it could also narrow the scope of searching for the murder suspect and direct the investigation.
PMSI (in water) unlike PMI (inland) is more difficult to estimate. Since the former is affected by additional factors (biotic and abiotic) such as more intense microbial metabolism, algal growth, and adipocere formation, it is more problematic.
In water, especially in cases of drowning, forensic practices increase in complexity. During drowning, water and microbes living in it rush into the respiratory and digestive tracts. This changes the endogenous microbiome and in turn, the inherent succession patterns (changes of the bacterial communities over time). Ruptures on the skin of the carcass further mess up previous succession patterns.
This is why most researchers so far have preferred to study death cases inland than on waters using PMSI through bacterial succession patterns. But now, many others are (fortunately) stepping up to investigate cases in water as well.
Microbial metabolism affects the rate of postmortem decomposition most, and this is why analyzing bacterial succession patterns over a longer time improves the accuracy of PMSI estimation.
Why employ bacteria in forensics?
We know that in the past forensic scientists heavily (or grossly) based their PMSI estimation on the science of forensic entomology – with the help of maggots crawling around the corpse, or the developmental stages of the fluttering flies near the corpse. However, these techniques have some setbacks of their own. These include other circumstances that hinder the investigation such as environmental factors influencing the growth stages of the insects, or the absence of such insects indoors or in winter.
For this reason, scientists have now turned to bacteria. They estimate the time elapsed since death by analyzing the stage of decay from bacterial decomposition.
How do you use bacteria to detect the time elapsed since death?
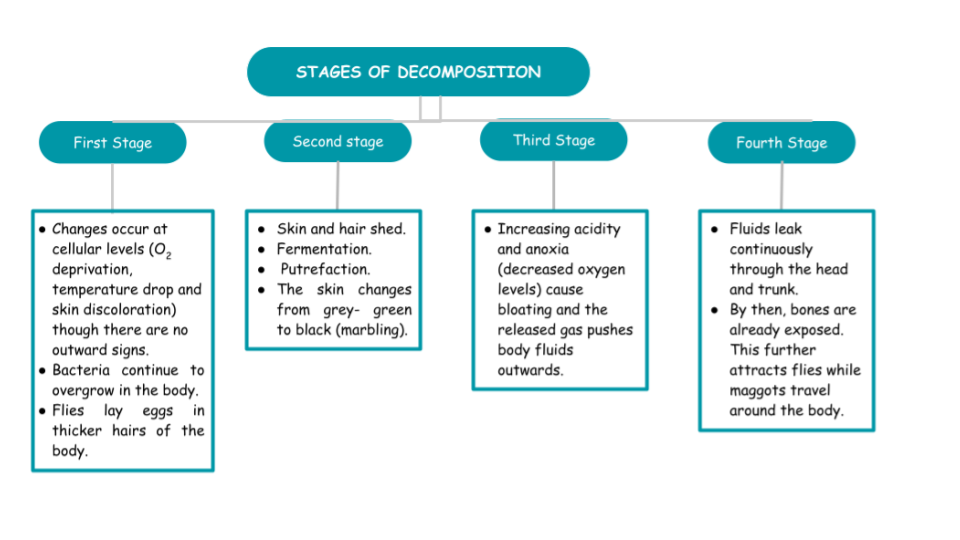
In her article, Hidaya Aliouche (2020) explains the stages of decomposition (Figure 2). She further states that different bacterial communities can indicate PMSI through bacterial succession that can be studied through high throughput sequencing (HTS). The data collected are then used to build machine learning algorithm-generated models based on the very temporal dynamic changes of the bacteria analyzed.
The study:
In 2022, Finkelbergs Dmitrijs and his research team investigated bacterial succession (community changes over time) in aquatic environments using epinecrotic biofilm on decomposing swine carcasses. Dr Jennifer L. Pechal defines epinecrotic bacteria as “those organisms residing, or moving, on the surface of decomposing remains (the skin or mouth)”.
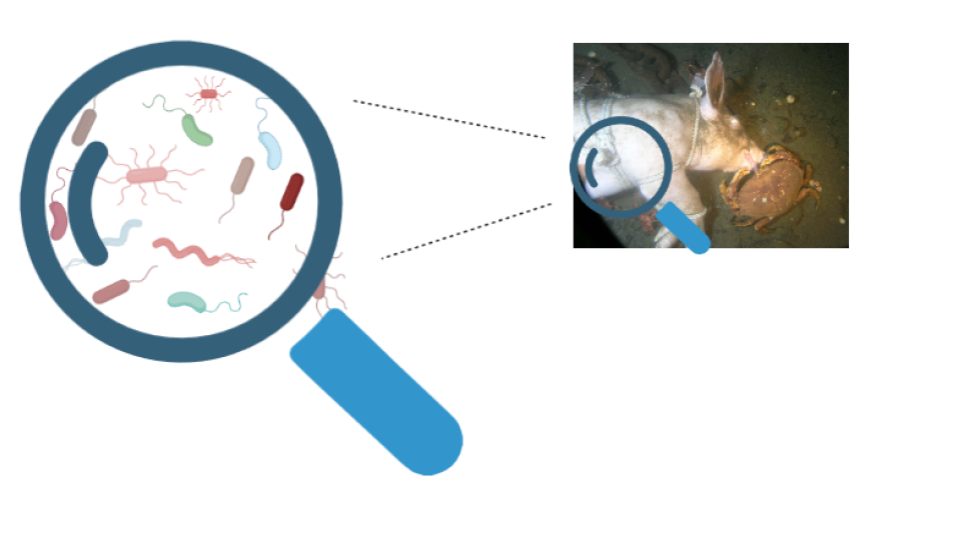
To identify the bacterial phyla present on the carcasses, the scientists sequenced the variable region 4 (V4) of 16S rDNA of the pigs’ carcasses, amounting to 131 samples in two trials: summer and winter. The total raw sequencing data was published in the Sequence Read Archive (SRA) under the accession number PRJNA841063 – feel free to check it out!
They found that the carcass decomposed much faster during summer (21 days) than in winter (56 days). The summer batch had twice as much diversity than the winter one! They concluded that seasonal variations indeed influenced the structure of the biofilm community on the carcass in water.
During the winter trial, they found that phyla like Firmicutes and Bacteroidetes present on the carcass surface were dominant, while during the summer, Firmicutes were mostly present in the epinecrotic community.
Moreover, the dominant bacteria in the epinecrotic (on the necrosis) community were foremostly heterotrophs and/or detritivores, while in the epilithic (on a rock surface) community they were predominantly autotrophs.
Taking the observations and deduced results in consideration, Dr F. Dmitrijs proposes implementing the succession patterns of epilithic biofilm coexisting with epinecrotic biofilm as a temporal control for PMSI estimation.
Perhaps, a better insight on the changing dynamics of bacteria during carcass decomposition?
Dr C. Hyun’s (2019) work is a good model to understand the bacterial dynamics during the various stages of decomposition. Dr F. Dmitrijs compares this with his own work to compare the bacterial communities to have an idea on the dynamics among the various phyla observed.
In Hyung’s study, Proteobacteria was the most dominant phylum which declined over time. Firmicutes started dominating as decay progressed. Bacteroidetes were detected only in the fresh and bloated stages. Actinobacteria found on the skin, which initially represented the majority of the communities, had long disappeared during the dry stage.
F. Dmitrijs’ study showed that Proteobacteria and Firmicutes were initially dominant – even before the body had entered the water. Proteobacteria dominated throughout the decomposition. Firmicutes, for their part, declined at the start but then picked up their abundance during the bloating stage. Bacteroidetes initially increased, then declined, and finally picked up until the carcass was in its sunken stage. Acidobacteria were opportunistically present once-a-while throughout the decomposition.
Though similar bacteria were observed in both works, their abundance varied considerably and contrastingly, depicting interesting trends. This comparison helps us to identify the key players (bacteria) and gives a way to look into the mystery behind the different trends as observed from the two studies.
Implications of the study and the door to future research:
Their work showed how bacterial dynamics and succession patterns changed with season and environmental factors. Based on this, they suggest that further research could be done to explore the effect of each environmental condition (temperature, light and water quality) on the rate of decomposition of the carcass.
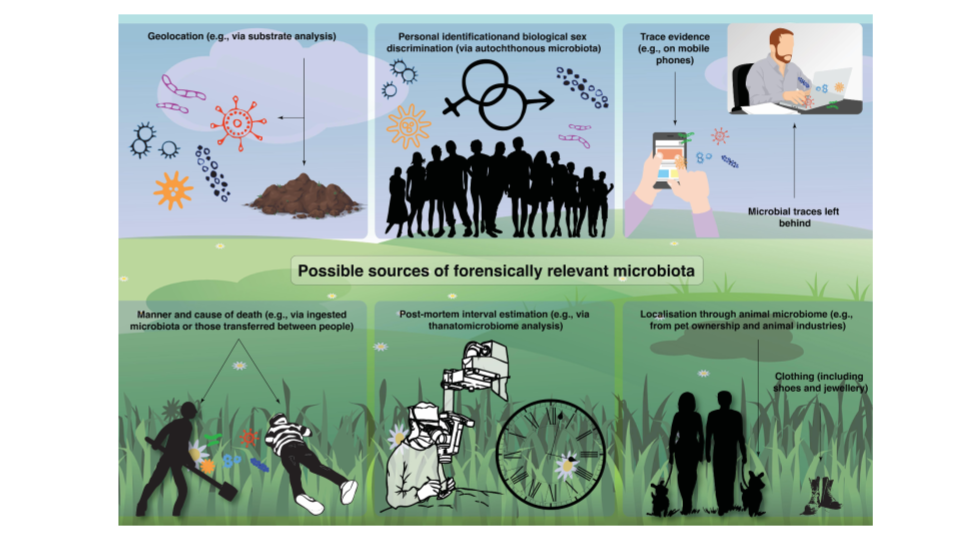
They also hope that further studies would widen the scope for forensic microbiology. This study, they claim, “provide[s] compelling evidence that the bacterial succession in the epinecrotic biofilm has a prominent potential to be used for the PMSI estimation in forensic investigations of submerged corpses.”
The author of this article (certainly) looks forward to seeing more forensic microbiology in action, what about you?
Link to the original post: Dmitrijs Finkelbergs et al (2022). Bacterial Succession in Microbial Biofilm as a Potential Indicator for Postmortem Submersion Interval Estimation. Frontiers in Microbiology; 13. Doi: https://doi.org/10.3389/fmicb.2022.951707
Additional references:
Featured image: Original image using shutterstock.com, icon-icons.com, dreamstime.com, thenounproject.com and biorender.com.



































































































