Breaking down the microbiology world one bite at a time
Soils: Sources or Sinks of Methane?
Our planet is warming fast because the greenhouse gases released into the atmosphere are trapping the heat. The most commonly known greenhouse gas is carbon dioxide (CO2) but there are others, including methane (CH4) and nitrous oxide (N2O).
In the atmosphere, methane is the second most abundant greenhouse gas (after carbon dioxide) but it is 28 times more potent than carbon dioxide! This is why emissions of methane, in particular, are closely monitored, especially in agricultural settings. For example, each year, a single cow can produce almost 100 Kg of methane by fermenting food in the rumen…That’s a lot of methane emitted by cattle each year!
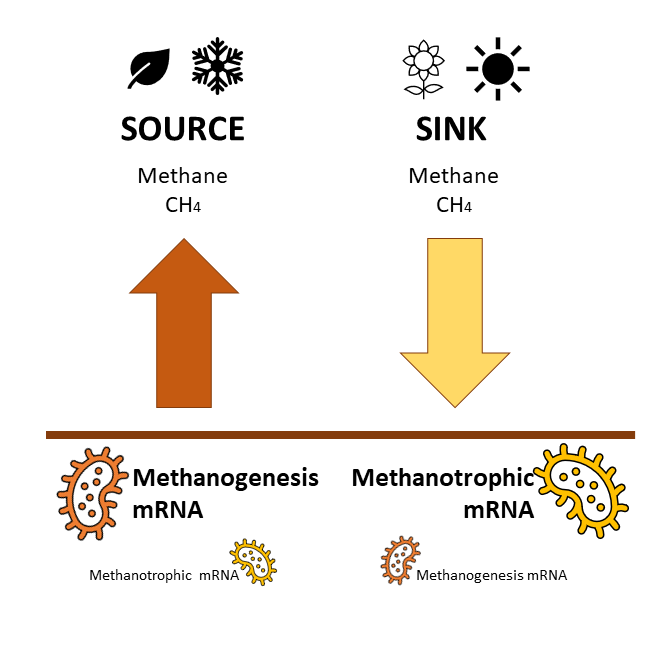
But, cows are not the only ones emitting methane into the atmosphere. Natural soils also produce methane, or we should say, microorganisms living in soils produce methane (methanogens); but some of them can also oxidise (degrade) that methane (methanotrophs). This fine balance between the abundance of these two types of microbes, methanogens and methanotrophs, determines whether soils could act as sources (emissions) or sinks (uptake) of methane. Understanding whether an ecosystem emits or uptakes methane can help model methane emissions and overall greenhouse gas emissions, but this is challenging…
While scientists can easily identify methanogens and methanotrophs using DNA-based omics methods, linking the diversity and abundance of these microorganisms to actual fluxes (emissions) of methane is quite a problem. To solve this, metatranscriptomics, an RNA- based omics method, has been employed. This allows the identification of active taxa (rRNA) and the quantification of active genes (using mRNA). This may, therefore, relate to measured fluxes (emissions) of greenhouse gases.
In this study, Täumer et al. (2022) used metatranscriptomics to relate methane cycling genes to methane fluxes and monitor seasonal changes in these fluxes. They sampled grassland soils in Germany during the autumn, winter, spring and summer seasons, and extracted the total RNA to be sequenced. They also measured emissions of methane and carbon dioxide at each sampling site using gas chambers placed in the soil.
They found that gas fluxes changed seasonally and that methane and carbon dioxide had surprisingly opposing trends. In autumn and winter, soils emitted methane and carbon dioxide emissions were low. During spring and summer, these soils, however, became sinks of methane but sources of carbon dioxide. These seasonally dynamic gas fluxes reflected changes not only in soil water content and temperature but also in the composition of the soil microbial community!
The team found a positive correlation between the abundance of methanogenesis mRNA transcripts and the methane fluxes measured in the field. In other words, more methanogenesis mRNA transcripts meant increased methane emissions.
In themselves, these results are a major step as they link omics’ methods to field observations. However, the team wanted to go further by identifying markers of methanogens and methanotrophs to determine if they could derive methane fluxes using metatranscriptomics. They showed that the ratio of methanogen/methanotroph (taxonomy using rRNA) could not be used to derive methane fluxes. However, the ratio of functional gene transcripts (methanogenesis/methanotrophic rRNA transcripts) reflects methane fluxes. This helps to determine whether soils are sources or sinks of methane.
Since soils form the major biological sinks of atmospheric methane (degraded by methanotrophs), monitoring changes in methane fluxes is essential, especially since emissions of greenhouse gases are increasing drastically globally. While it is still difficult to link specific biological communities to gas fluxes, this study brings us one step closer to understanding methane fluxes.
Link to the original post: Täumer, J., Marhan, S., Groß, V., Jensen, C., Kuss, A. W., Kolb, S., & Urich, T. (2022). Linking transcriptional dynamics of CH4-cycling grassland soil microbiomes to seasonal gas fluxes. The ISME Journal, 1-10.
Additional sources: Soils as sources and sinks for atmospheric methane
Featured image: