
Breaking down the microbiology world one bite at a time
There’s no such thing as inefficiency.
Nitrogen is the main constituent of all life, being essential for e.g. proteins and DNA. Despite its abundance on earth, nitrogen supply is a growth-limiting factor for most life forms. I’m not talking about fitness enthusiasts chugging protein shakes for muscle mass, but – of course – about microorganisms. Bacteria and archaea are the only life forms capable of fixing nitrogen out of the air, and into cells. That means all nitrogen in our bodies (but also in that of other animals or plants) has been “prepared” for us by bacteria and archaea. The only enzymes known to catalyze such a process are called nitrogenases.
Three variants of these nitrogenases exist, named after different metals found in their active site: molybdenum (Mo), vanadium (V) and iron (Fe). The enzymes take nitrogen from the atmosphere and some source of electrons to reduce N2 to NH3 but also produce a range of byproducts (chiefly H2). Of the three variants, Mo-nitrogenase is the oldest enzyme and the other two evolved from it (1). However, Mo-nitrogenase is also considered the most efficient nitrogen fixation enzyme because it produces the least by-products (1). So if the V- and Fe-nitrogenase are less efficient at their job, why would they have evolved at all?
Katja Luxem and colleagues at Princeton University, NJ, set out to find answers using Rhodopseudomonas palustris (2). This bacterium contains the genes for all three enzymes, but – as the article shows – usually depends on Mo-nitrogenase for its nitrogen supply. In addition, the researchers tested three genetic mutants of R. palustris which each contained only one of the nitrogenase isozymes. They found that, depending on the carbon source, Mo-nitrogenase did not fix nitrogen the fastest. V-nitrogenase supported faster growth on other substrates, and on acetate even outperformed the wild-type (Figure 1).
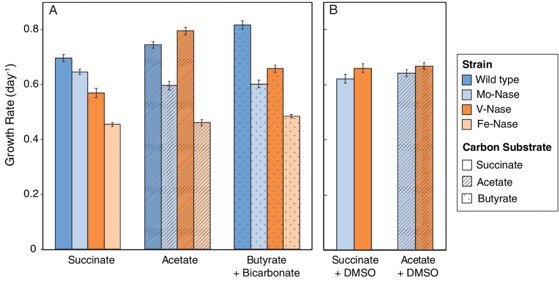
When fed with acetate, the mutant expressing only V-nitrogenase produced about as much byproduct (H2) as the wild-type. The authors suspect it has to do with temperature: the idea that Mo-nitrogenase is the most efficient arose from experiments at 30 °C, but this experiment was carried out at 19 °C – and they’re not the first to see the efficiency of the Mo nitrogenase drop at lower temperatures (3). The existence of the V-nitrogenase next to the Mo-nitrogenase might therefore have to do with the temperature of the environment that the bacterium lives in.
Luxem and colleagues also show that nitrogen fixation efficiency does not solely depend on the nitrogenases, but on a whole cascade of enzymes and cofactors delivering the electrons and energy for the catalysis. Using proteomics they found that the V-nitrogenase mutant, which performed so well on acetate, was backed up by a system of energy delivery proteins very similar to the wild-type. The mutant carrying only the Mo-nitrogenase had a different profile of supporting enzymes. The evolution of different nitrogenase forms might therefore also be linked to the interplay with other proteins.

Interestingly, R. palustris does not compensate for slower nitrogenases by spending more resources on nitrogen fixation. Apart from nitrogen, R. palustris can also fix carbon from CO2 (in a process similar to photosynthesis). To fix either element, electrons are required. The bacterium therefore has to balance how much of each it fixes (see Figure 2, arrows E and F). In Luxem’s experiments, nitrogen supply is growth-limiting to R. palustris (and is given all the carbon source it could need). It would therefore seem useful to redirect as many electrons as possible to nitrogen fixation. However, the authors report something different: the percentage of available electrons used for N-reduction stays constant, around 60%, for different carbon sources and nitrogenase enzymes. This is especially surprising for the Fe-nitrogenase, which fixes very little N2 at the expense of producing a lot of H2. The authors propose that some (yet unknown) evolutionary selection pressure exists against abolishing carbon fixation.
In sum, it seems that the “useless” V-nitrogenase was simply never examined in a way that showed its strength: nitrogen fixation at lower temperatures. Thanks to Luxem and her team, we have seen a glimpse of what it’s good at. And in due time we’ll undoubtedly get to appreciate the advantage of Fe-nitrogenase, too!
Link to the original post: Carbon substrate re-orders relative growth of a bacterium using Mo-, V-, or Fe-nitrogenase for nitrogen fixation Katja E. Luxem, Anne M. L. Kraepiel, Lichun Zhang, Jacob R. Waldbauer, Xinning Zhang
Featured image: https://commons.wikimedia.org/wiki/File:Nitrogen_Cycle_1.svg
Written by Steven Pilon



































































































