Breaking down the microbiology world one bite at a time
A new letter in the genetic alphabet.
All DNA is made using the same alphabet. From viruses to human beings, we all use the four letter code: A, C, T, G. These four nucleotides (Adenine, Cytosine, Thymine and Guanine) make out the genetic code of all organisms.
However, in 1977 an anomaly was found. A bacteriophage (S-2L) that infects cyanobacteria was found to not have the letter A but a chemically-related nucleotide: diaminopurine or Z. DNA modifications are not uncommon, but here, the new letter completely replaced A in the phage DNA pairing with its complementary base T, while C still paired with G.

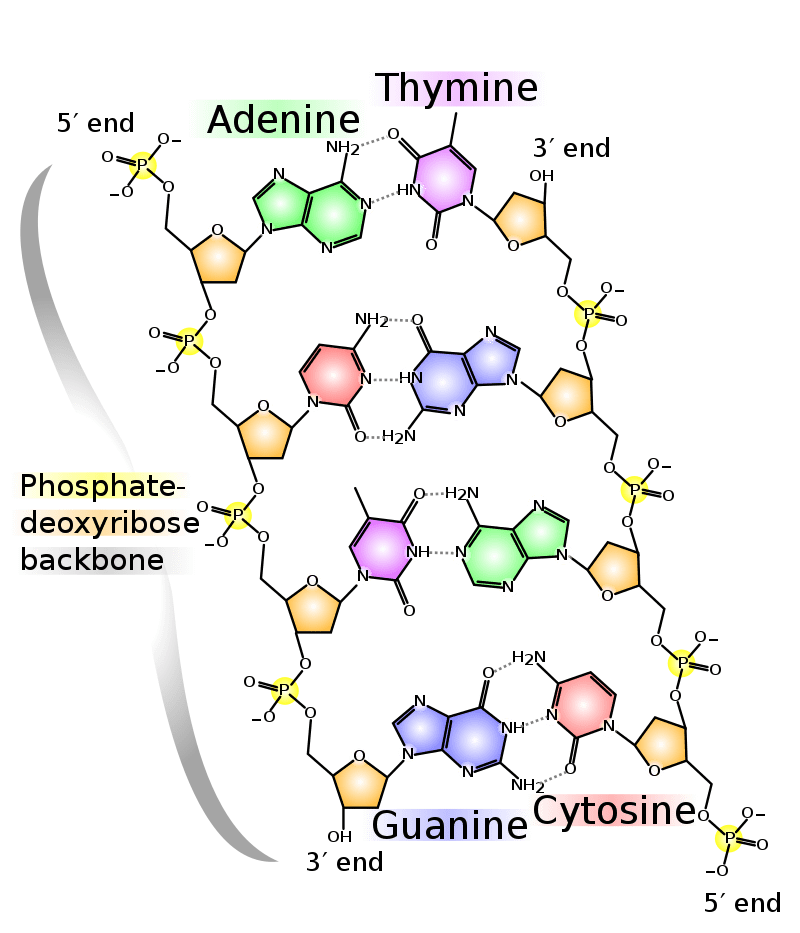
Since then, other studies have looked into this new DNA letter in terms of synthetic pathways and structure within the DNA, but with limited results. The DNA (deoxyribonucleic acid) is composed of a backbone made of alternating sugar-phosphate arrangement. These phosphates give the DNA its acidic properties, while the sugar is a pentose called 2-deoxyribose. The nucleic part of the name comes from our four letter code, the nucleotides. The nucleotides keep the two strands in a double helix by a force of attraction consisting of hydrogen bonds. The A-T pair forms two hydrogen bonds, while C-G forms three, as seen on the figure.
In 1998, a study showed that the Z-T pair in the S-2L lytic phage was forming three hydrogen bonds, thereby changing the 3-D structure of the DNA (more about lytic phages here).

However, it wasn’t until April 2021 that several scientific groups from France and China found the same “anomaly” in other bacteriophages, and were able to better characterize the mechanisms of the new biosynthesis pathway as well, as its implications (REF1, REF2, REF3).
Both Yan Zhou’s and Dona Sleiman’s research groups showed that the Z letter was used by other phages, while the third study was focused on a Vibrio phage Phi-VC8. By studying how the new nucleotide is used in the different organisms, both Zhou and Sleiman’s groups looked at the synthetic pathways involved in the synthesis of Z and the enzyme PurZ in more detail.
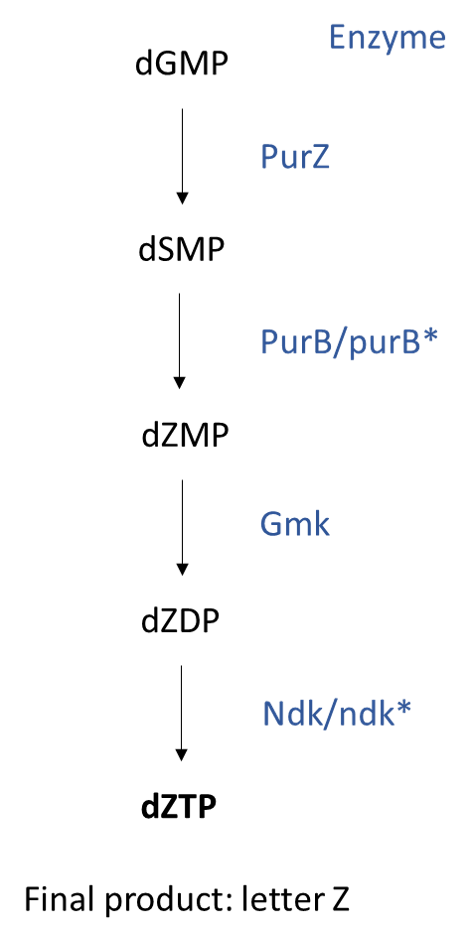
Using several mutants, Sleiman and colleagues showed which enzymes were essential to synthesize the new nucleotide, and were able to describe its synthesis from dGMP (guanine monophosphate) to dZTP (Z triphosphate). They showed that purB* mutants had a lower infection efficiency, while ndk* mutants did not infect cells at all. These results highlight that these genes are needed for dZTP synthesis, and without them, phages cannot replicate.
The latest study of Valerie Pezo and colleagues focused more on the DNA replication machinery, specifically on the DNA polymerase (more about DNA machinery here). DNA polymerases are used to replicate DNA: they bind to each single stranded side and read the sequence in a 5’-3’ way. As it reads, it adds the complementary nucleotides (T for A, C for G and vice versa) from their free NTP form (nucleotide triphosphate, TTP, ATP, GTP or CTP).

Wikicommons
The question is: do these phages have a different DNA polymerase to incorporate ZTP when replicating their DNA? The study showed that the DNA polymerase found in organisms that use ZTPs were from 30 to 90 times more efficient at using ZTPs instead of ATPs. Conversely, E. coli’s DNA polymerase was about twice more efficient inserting ATP when both ATP and ZTP were put in the cell. Therefore, it seems that these organisms do have a DNA polymerase optimized to use the new alphabet.
What are the implications? Why did these viruses evolve this way? Zhou and colleagues believe it gives an evolutionary advantage to the phages by letting them evade host defences, carried-out by restriction enzyme attacks. Understanding this new alphabet could have a major impact on different fields such as making nanostructures from DNA, also called DNA origami, or in DNA-based data storage. Finally, this new alphabet could be used to engineer phages to increase host cell range and efficacy of phage therapy, as well as food preservation among other potential uses (read more about phage therapy here).
Maybe the DNA alphabet we know today will expand as we study more and more organisms, especially viruses. While we have engineered new letters ourselves, it seems that nature already did the job for us.
Link to the original post:
Featured image: Modified from: https://innovativegenomics.org/glossary/dna/