Breaking down the microbiology world one bite at a time
Moving across borders: how bacterial genes become mobile
Bacteria carry different construction manuals within their DNA to build enzymes or other proteins which enable the movement of DNA within their own genome or between different bacteria [1]. Those construction manuals are also called mobile genetic elements (MGEs) and include for example plasmids, bacteriophages, and transposons. Whereas plasmids are organized into stable, self-replicating entities or “replicons” within the bacteria, bacteriophages can inject their genome and integrate it into the bacterial cell and chromosome. On the other hand, transposons can randomly “hop” into phage or plasmid DNA which potentially changes the properties of the host cell [1]. All these mechanisms allow certain genes to move from one bacterial cell to the next, and on a larger scale to travel from one habitat such as the human body to the next, for example, animals or the environment.
Research suggested the presence of antibiotic resistance genes (ARGs) within MGEs. Carbapenem is a class of antibiotics with a broad spectrum of antimicrobial activity against gram-positive and gram-negative bacteria. One mechanism by which these bacteria gain resistance against carbapenem is the expression of carbapenemases. Carbapenemases belong to the group of hydrolases that hydrolyze and therefore inactivate antibiotics of the Carbapenem class. Carbapenemase-encoding genes such as blaOXA-48 are found in plasmids of Klebsiella pneumoniae where they are responsible for the expression of Carbapenemases [2]. Carbapenem-resistance Enterobacterales are a group of bacteria including Klebsiella pneumoniae, Acinetobacter baumannii, or Pseudomonas aeruginosa, that commonly cause infectious diseases in healthcare settings e.g. hospitals, but for which treatment with carbapenem antibiotics fail [3]. Even though this group of bacteria has been extensively studied in humans, there is little known about the epidemiology and transmission of them in the environment or animals.
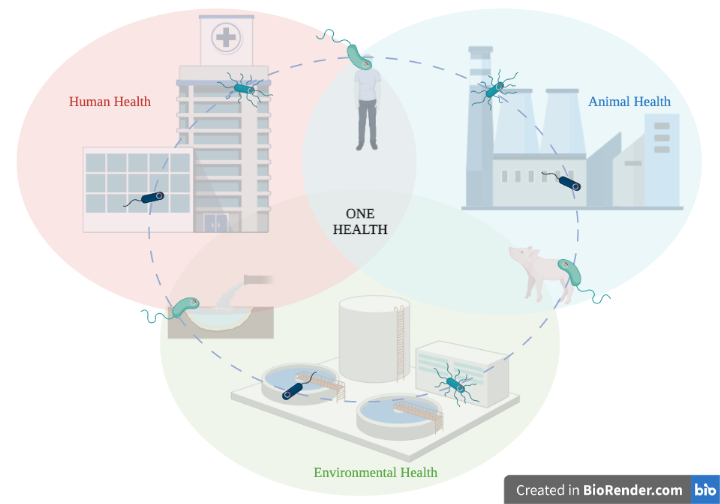
Humans are constantly in contact with bacteria through their environment for example by touching surfaces, plants, animals, or by eating and drinking. These are all travel routes for bacteria and their antibiotic resistance genes to move across human borders and enable them to spread not only on a local but also global level. Thus, researchers and health organizations are now focusing on a so-called “One Health” approach to combat the spread of antimicrobial resistance, which focuses on collaborative action across the human, environmental, and animal sectors [4] (Figure 1).
Ramsamy and his colleagues have studied and compared MGEs of Carbapenem-resistance Enterobacterales in humans, the environment, and animals within the same geographical area in South Africa using a “One Health” approach [5]. The researchers were able to isolate nineteen carbapenemase-producing Enterobacterales from three public sector hospitals, and waterbodies from a wastewater treatment plant, and found a high prevalence of K. penumoniae (47%) and Enterobacter hormaechei (32%). However, they found no isolates from pigs at a food processing plant with a slaughterhouse.
Afterward, they characterized the isolates as multidrug-resistant to antibiotics such as ceftazidime, cefuroxime, and ertapenem. Additionally, sixteen out of nineteen isolates that were resistant against at least one carbapenem also contained carbapenemase-encoding genes with the highest prevalence for blaOXA-181 and blaNDM-1. Both genes were common in human as well as environmental isolates.
In total, the research group identified forty-two plasmids from the nineteen isolates. Within the plasmids, they found ARGs together with transposable elements and/ or insertion sequences. For example, a closer look at the genetic environment surrounding the Carbapenemase-encoding gene blaOXA-181 revealed the presence of the ereA gene. The latter encodes for erythromycin esterase, which conveys high resistance to the antibiotic erythromycin. Moreover, insertion sequences such as IS3000 and transposons such as Tn3 were also found in plasmids that harbor blaOXA-181. Thus, one can imagine MGEs as a vehicle that takes several antibiotic resistance genes on board and moves between bacterial cells into new sectors.
However, the researchers also pointed out the limits of this study in terms of generalizability because of the small number of isolates obtained. To confirm the insights gained about the spread of antimicrobial resistance in a “One Health” context, they suggested further research with larger sample sizes and in different geographical locations.
Currently, antibiotic resistance genes have been found on MGEs in high arctic soil [6], as well as drinking water in India [7]. This indicates the far-reaching spread of antibiotic resistance to different sectors around the world and suggests MGEs as a driver. However, MGEs can carry not only antibiotic resistance genes but also virulence or other resistance genes which implies further consequences on human health for existing bacterial infectious diseases [3].
Link to the original post: Ramsamy, Y., et al. (2021). “Mobile genetic elements-mediated Enterobacterales-associated carbapenemase antibiotic resistance genes propagation between the environment and humans: A One Health South African study.” Sci Total Environ 806(Pt 3): 150641.
Featured image: Photo by Norbert Dudas on Unsplash
References:
- Frost, L. S., et al. (2005). “Mobile genetic elements: the agents of open source evolution.” Nat Rev Microbiol 3(9): 722-732.
- Yang, X., et al. (2021). “Carbapenem Resistance-Encoding and Virulence-Encoding Conjugative Plasmids in Klebsiella pneumoniae.” Trends Microbiol 29(1): 65-83.
- CDC – Carbapenem-resistant Enterobacterales (CRE) – https://www.cdc.gov/hai/organisms/cre/index.html November 9, 2021
- WHO – WHO,FAO, and OIE unite in the fight against Antimicrobial Resistance – https://www.who.int/foodsafety/areas_work/antimicrobial-resistance/amr_tripartite_flyer.pdf November 9, 2021
- Ramsamy, Y., et al. (2021). “Mobile genetic elements-mediated Enterobacterales-associated carbapenemase antibiotic resistance genes propagation between the environment and humans: A One Health South African study.” Sci Total Environ 806(Pt 3): 150641.
- McCann, C. M., et al. (2019). “Understanding drivers of antibiotic resistance genes in High Arctic soil ecosystems.” Environ Int 125: 497-504.
- Walsh, T. R., et al. (2011). “Dissemination of NDM-1 positive bacteria in the New Delhi environment and its implications for human health: an environmental point prevalence study.” Lancet Infect Dis 11(5): 355-362.