Breaking down the microbiology world one bite at a time
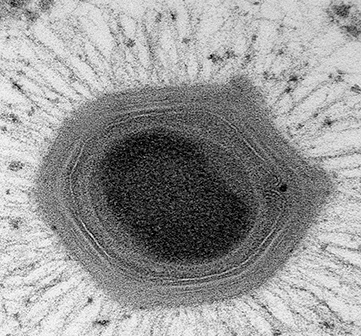
Mimivirus – a giant in a small world.
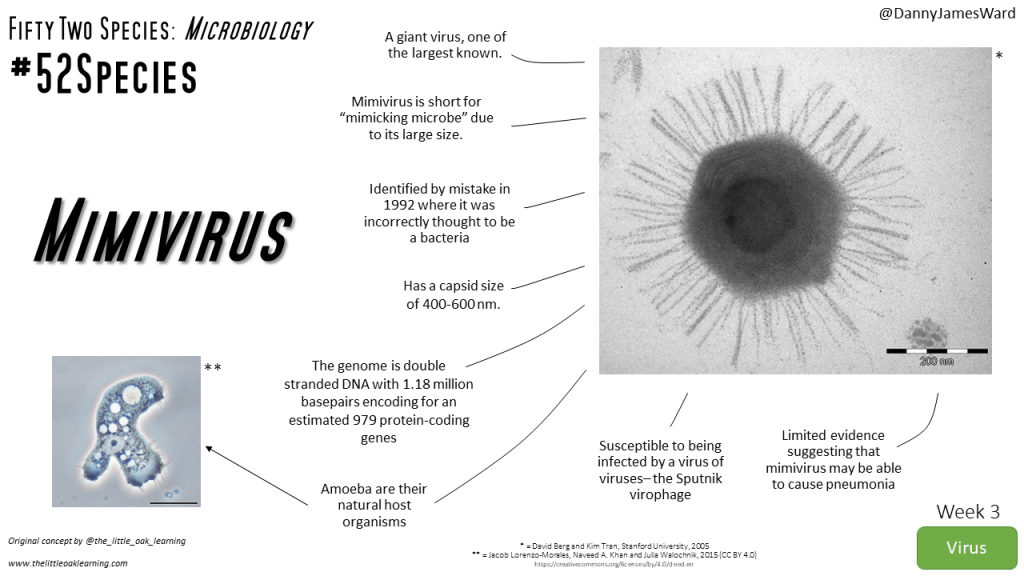
Mimiviruses are part of the largest known viruses: 2 to 40 times bigger than other viruses. They are so big they were found to be infected by smaller viruses called Sputnik virophages. Mimiviruses were discovered in 1992 and were thought to be bacteria based on preliminary observations and therefore were called Mimivirus from “mimicking microbes”. These viruses are found in amoebas, which are unicellular organisms.


To learn more about the different types of viruses and their different sizes and shapes visit National Geographic:
The origin of giant viruses is still not well known. However several hypotheses have been proposed:
- The viral world existed before the LUCA (Last Universal Common Ancestor – Forterre and Prangishvili 2013)
- Giant viruses evolved from their cellular hosts by reductive evolution – organisms that are losing genes (Claverie 2006; Nasir et al. 2012a; Nasir and Caetano-Anollés 2015)
- Giant viruses evolved by host gene capture and gene duplication events. Organisms that had a small genome but expended their genome by acquiring host genes or duplicating genes (Shackelton and Holmes 2004; Moreira and Brochier-Armanet 2008)
- Giant viruses originated from small viruses and acquired genes from eukaryotic hosts and other cellular organisms by horizontal gene transfer (transfer of DNA between organisms) (Koonin, Krupovic, et al. 2015; Koonin and Yutin 2018).


Other groups are proposing to reclassify the giant viruses in a fourth domain as TRUC (Things Resisting Uncompleted Classification) (Boyer et al. 2010; Raoult 2013)
To understand the origin of giant viruses, Patil and Kondabagil looked at the complexity and the evolution of the Mimivirus DNA replication machinery. To do this, they compared the machinery between different organisms: another virus – bacteriophages (T4), bacteria (E. coli), archaea, eukaryotes, and Mimivirus.
DNA replication is a complex process involving many proteins to open the double DNA strands, to remove the supercoil ahead of the replication fork, to make the primer and remove them, to copy the DNA, to link the DNA fragments, etc… (Figure and video below)

The different types of proteins involved are different for each type of organism compared in this study. In T4 bacteriophages, the machinery is one of the simplest ones, and as complexity increases with genome size, the E. coli machinery involves more proteins (about 30). In archaea, the machinery has processes similar to both bacteria and eukaryotes. The system itself is comparable to eukaryotes while the proteins are closely related to bacteria. Finally, in eukaryotes, the process is even more complex as the genomes are much larger, and therefore requires more proteins.
The authors compared the machinery between the different organisms and concluded that the Mimivirus replication machinery came early in the evolution of the virus, supporting the hypothesis of Mimivirus evolution from a complex ancestor.
But which ancestor? Well, two hypotheses remain:
- From a complex cellular ancestor by losing genes (hypothesis 2)
- From a simple viral ancestor which gained complexity by acquisition and accumulation of genes from external organisms (hypothesis 4)

The authors showed that out of the nine replication machinery proteins investigated, five are of eukaryotic origin, two are of bacterial, one of phage origin, and one protein with unknown origin.
Based on these results, they propose that Mimiviruses might have evolved from a complex cellular ancestor that over time lost genes by reductive evolution (hypothesis 2).

However, from which ancestors these giant viruses originate is still not understood and more studies on giant viruses will be needed to fill in this gap of knowledge. But for now, the authors suggest that hypothesis 2 is the hypothesis most likely to explain the origin of giant viruses including Mimivirus.
Learn more about Mimivirus:
Link to the original post: Supriya Patil, Kiran Kondabagil, Coevolutionary and Phylogenetic Analysis of Mimiviral Replication Machinery Suggest the Cellular Origin of Mimiviruses, Molecular Biology and Evolution, 2021
Doi: https://doi.org/10.1093/molbev/msab003